In real-time quantitative PCR (qPCR) experiments, it is not uncommon to encounter weak probe signals, significant Ct value variability, or noticeably lower signal intensity in one channel during fourplex detection.These issues are often not caused by the template or primers, but are closely related to probe molecular design.
This article uses the optimization of Texas Red probes as an example to systematically review key aspects of probe design, including quencher selection, dye-instrument compatibility, and comprehensive validation of four-color systems. Through this case study, we aim to help identify potential design pitfalls and demonstrate with experimental data that optimization truly makes a difference.
How TaqMan Probes Work
A typical TaqMan probe contains a reporter dye at the 5′ end and a quencher at the 3′ end.
When the probe is intact, the reporter and quencher are in close proximity, suppressing fluorescence and maintaining a low background. During PCR extension, Taq polymerase cleaves the probe, separating the reporter from the quencher and allowing fluorescence to be emitted.
The stronger the signal and the lower the background, the more accurate the quantification.
While the mechanism is simple, achieving optimal probe performance in real experiments is not always straightforward.

Why Do Probes Sometimes Produce Weak Signals
1. Distance Between Reporter and Quencher
An ideal quencher should have a broad absorption spectrum and high quenching efficiency. However, in short probes less than 20 nucleotides, the physical distance between the 5’ reporter and 3′ quencher may be insufficient for complete quenching, resulting in elevated background fluorescence.
Introducing internal quenchers can help reduce this distance, but may interfere with the 5′ to 3′ exonuclease activity of Taq polymerase, reducing cleavage efficiency and signal release.
2. Dye Compatibility with qPCR Instruments
Different qPCR platforms have distinct optical detection systems, making dye selection critical.
For example, on the Roche LightCycler 480 platform, detection efficiency varies significantly between dyes. Choosing an incompatible dye can directly lead to reduced signal intensity.
3. Channel Interference in Multiplex qPCR
Fourplex qPCR using FAM, SUN, Texas Red, and Cy5 requires minimal spectral overlap and consistent amplification efficiency across all channels.
Poor probe performance in any one channel can compromise the stability and reliability of the entire assay.
Case Study: Optimizing Texas Red Probes
To address weak signals in short Texas Red probes of approximately 18 nucleotides, we performed a systematic optimization of quencher combinations.
By adjusting both the type and placement of quenchers, we improved the spatial relationship and energy transfer efficiency between the reporter dye and quencher, while preserving Taq polymerase activity.
The optimized probes maintained low background fluorescence and significantly increased signal intensity, with clear improvements in endpoint RFU.
Additionally, selecting reporter dyes better matched to the Roche LightCycler 480 system further enhanced signal performance.
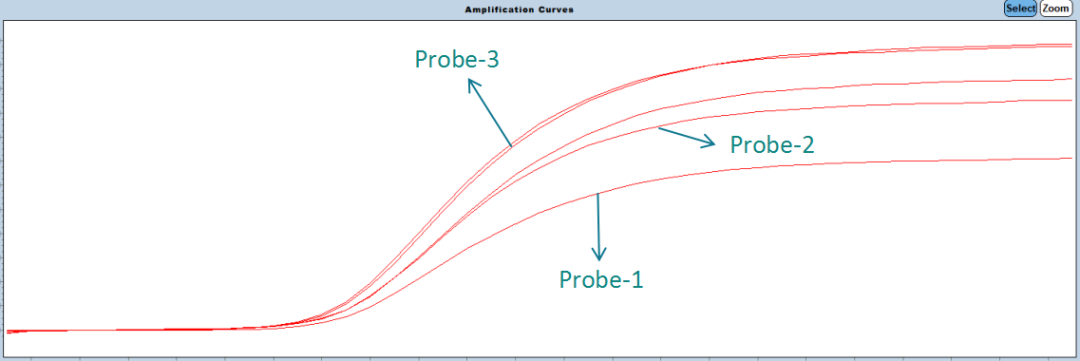
Probe-3 : After the quenching group is optimized, the signal intensity is significantly improved.
Head-to-Head Comparison with a Competitor
Based on these optimization strategies, we designed a set of qPCR probes using FAM, SUN, Texas Red, and Cy5, and compared them with probes of identical sequences from another supplier.
All experimental conditions were identical, including primers, template, and instrument. The only difference was the probe source.
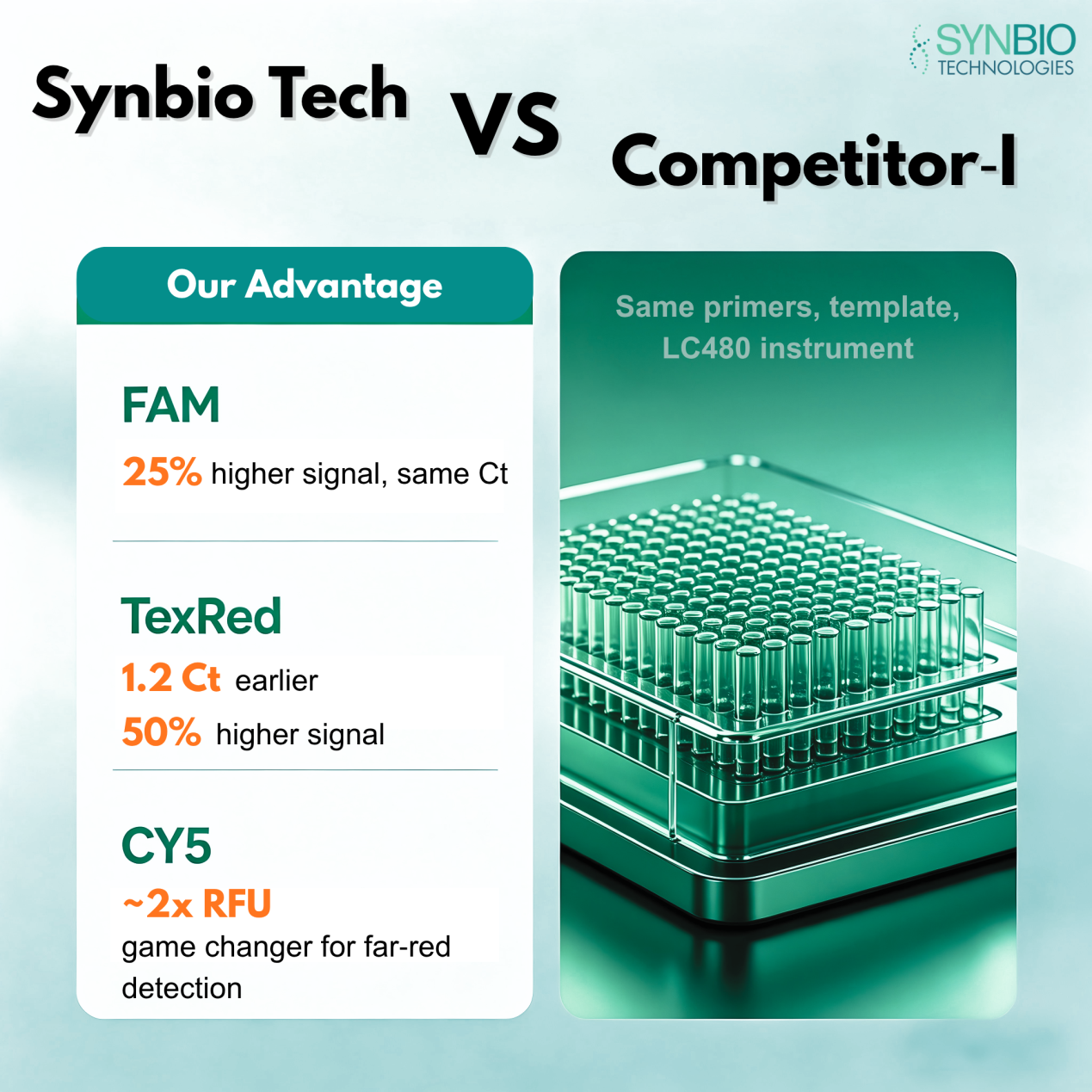

Results Overview
-
FAM and SUN Channels
Ct values were similar, but our FAM probes showed a 25 percent higher signal intensity, indicating improved sensitivity and signal-to-noise ratio.
-
Texas Red Channel
Our probes achieved a 1.2 cycle Ct advantage and a 50 percent increase in signal intensity. For low-copy samples, this difference can determine whether a signal is detectable.
-
Cy5 Channel
RFU values nearly doubled, effectively addressing the common issue of weak far-red signals, which is critical in multiplex assays.
Synbio Technologies: What Makes a High-Performance Probe
1. Data-Driven Modification Strategies
Effective probe design is not about adding more modifications, but about selecting the right ones based on experimental validation. The goal is to balance sensitivity, specificity, and signal-to-noise ratio.
2. Instrument-Specific Optimization
Different qPCR platforms have unique spectral responses. Matching probe dyes to the instrument improves detection efficiency and overall performance.
3. Balanced Performance in Multiplex Systems
Multiplex qPCR requires not only strong individual channel performance, but also stable coordination across all channels. Optimization in challenging channels such as Texas Red and Cy5 is especially important.
4. Technical Support Matters
Probe design challenges and unexpected signal issues are common. Professional technical support can help identify root causes and provide effective optimization strategies from design to application.
Conclusion
A high-quality probe is not defined by complexity, but by precision and validation.
Even small design improvements can lead to significant gains in signal strength, accuracy, and reproducibility. In qPCR, details matter, and optimized probe design makes all the difference.
 DNA Synthesis
DNA Synthesis Vector Selection
Vector Selection Molecular Biology
Molecular Biology Oligo Synthesis
Oligo Synthesis RNA Synthesis
RNA Synthesis Variant Libraries
Variant Libraries Genome KO Library
Genome KO Library Oligo Pools
Oligo Pools Virus Packaging
Virus Packaging Gene Editing
Gene Editing Protein Expression
Protein Expression Antibody Services
Antibody Services Peptide Services
Peptide Services DNA Data Storage
DNA Data Storage Standard Oligo
Standard Oligo Standard Genome KO Libraries
Standard Genome KO Libraries Standard Genome Editing Plasmid
Standard Genome Editing Plasmid ProXpress
ProXpress Protein Products
Protein Products