Genome-wide knockout screening has become a core technology in functional genomics, drug target discovery, and disease mechanism research. At the center of every successful screening experiment lies the genome-wide sgRNA library system, whose quality directly determines the accuracy, reproducibility, and robustness of downstream functional readouts.
The general workflow of gene knockout screening is well established: a genome-wide library containing tens of thousands of guide RNAs is delivered into target cells via lentivirus transduction, followed by application of selective pressure. Next-generation sequencing (NGS) is then used to quantify enrichment or depletion of guide RNAs, enabling the identification of genes associated with a specific phenotype.
While the conceptual framework appears straightforward, one critical factor is often underestimated: the initial library representation and distribution bias have a direct impact on the reliability of screening outcomes.
Leveraging its proprietary library construction platform and high-throughput chip synthesis technology, Synbio Technologies has developed a fully integrated workflow solution for genome-wide knockout screening projects. Using a mouse genome-wide knockout library and downstream cell pool construction as a representative example, this case study highlights how a end-to-end service model ensures process consistency, data traceability, and experimental reproducibility from start to finish.
Project Background and Experimental Requirements
The client was a cancer research group focused on identifying genetic drivers of tumor drug resistance using a genome-wide functional screening approach.
The project required a fully integrated workflow covering library construction, lentivirus packaging, and cell pool construction, with the following specifications:
Library Construction Requirements
-
High-purity, endotoxin-free plasmid preparation
-
Library representation >99%
-
Distribution uniformity <10
Lentivirus Production Requirements
-
lentivirus vector system for stable gene perturbation
-
Viral titer ≥1 × 10⁸ TU/mL
-
Sterility and mycoplasma-free confirmation
CelPoolConstruction Requirements
-
Cell line: L929 mouse cells stably expressing Cas9-BSD
-
Culture medium: DMEM + 10% FBS
-
Adherent culture system
-
Passage ratio: 1:5, ~48-hour confluency cycle
Selection Validation
-
Puromycin concentration: 10 μg/mL
-
Complete elimination of non-transduced cells within 48 hours
NGS Performance Criteria
-
Library coverage >85%
-
Distribution uniformity <15
Service Model Requirement
-
Fully integrated, one-stop delivery
-
No intermediate client handling required
-
End-to-end traceability and process control
-
Reduced experimental turnaround time
Workflow Implementation and Results
The project was executed through a fully integrated three-stage workflow:
1. Genome-wide knockout library plasmid preparation
2. lentivirus production and validation
3. Celpooconstruction
All stages were managed under standardized operating procedures with closed-loop quality control.
I. Library Plasmid Construction: Ensuring High-Fidelity Genome Coverage
The plasmid library serves as the foundational input for all downstream screening steps. Any bias introduced at this stage will propagate through viral production and cellular selection.
An endotoxin-free maxiprep workflow was combined with NGS-based quality validation.
Results demonstrated:
-
Library representation: 99.95%
-
Distribution uniformity: 4.0
These results indicate highly even guide RNA distribution with minimal amplification bias, ensuring robust genome-wide coverage and high confidence in downstream functional screening.
II.lentivirus Production: Enabling Efficient and Stable Gene Perturbation
lentivirus production is a critical step that bridges the plasmid library and functional cellular screening system. Viral titer and infectivity directly determine transduction efficiency and the likelihood of achieving single-copy genomic integration.
To validate viral functionality and optimize antibiotic selection conditions, a pilot transduction experiment was conducted using L929 Cas9-expressing cells.
After 48 hours of viral exposure, cells were subjected to puromycin selection (10 μg/mL).
Observations (40×):
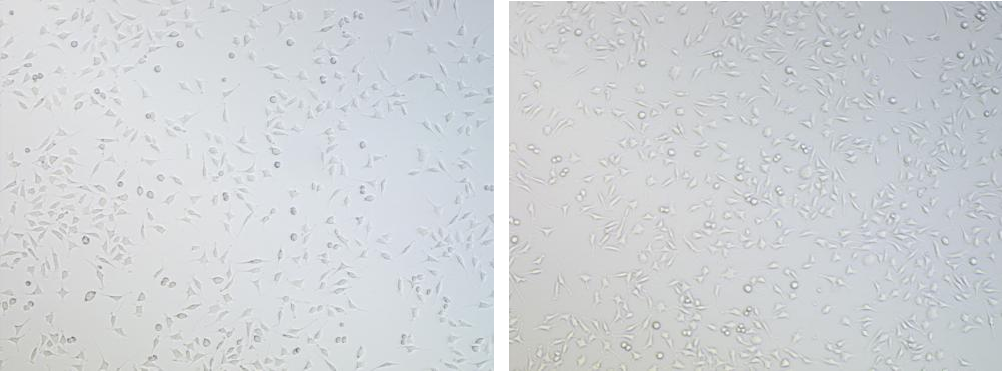
Pre-screening control group Pre-screening test group
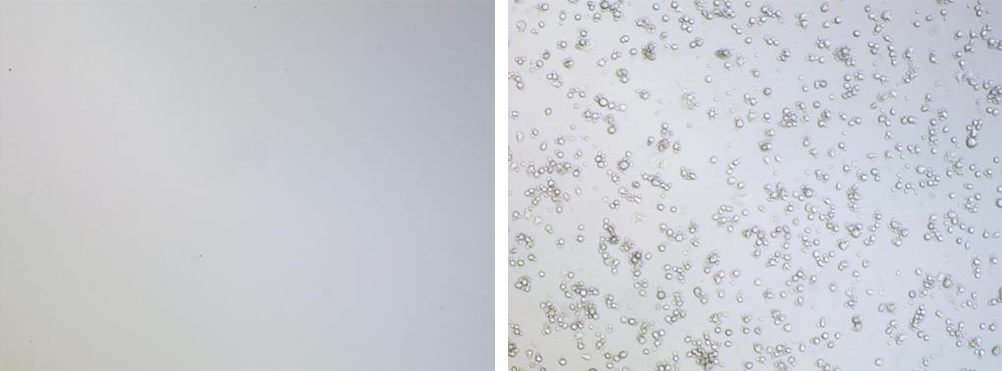
Post-screening control grou Post-screenin test group
-
Non-transduced control cells were completely eliminated within 48 hours
-
Transduced populations retained robust viable colonies with stable adherence
Based on this validation, a puromycin concentration of 10 μg/mL for 2 days was adopted for the full-scale workflow.
III. sgRNA Library Cell PooConstruction
The sgRNAlibrarycellpool represents the functional execution stage of genome-wide screening, where library representation is converted into a measurable cellular phenotype.
Key quality parameters include transduction efficiency, single-integration probability, and maintenance of library diversity during cell expansion.
Final results showed:
√ Library coverage: 97.93%
√ Distribution uniformity: 8.8
These metrics confirm strong preservation of library complexity throughout viral delivery and cell expansion, ensuring reliable downstream screening performance.
The resulting celpool population was delivered as a screening-ready system, requiring no additional processing from the client and enabling immediate use in functional genomic assays.
Conclusion and Key Insights
This case demonstrates the successful implementation of a fully integrated genome-wide knockout screening workflow, including:
1. High-quality library construction
2. lentivirus production and functional validation
3. Cellpooconstruction
All performance metrics exceeded client requirements, providing a reliable foundation for identifying genes associated with tumor drug resistance.
More importantly, this study highlights the value of a fully integrated service model. By unifying library preparation, viral production, and cellular construction into a single controlled workflow, this approach significantly reduces technical variability and improves reproducibility.
For researchers, the advantages are clear:
√ Improved experimental consistency
√ Reduced workflow complexity
√ Shorter project timelines
√ Enhanced data reliability
√ Greater focus on biological interpretation rather than process management
As genome-wide functional screening continues to expand across oncology, immunology, and drug discovery, integrated end-to-end workflow solutions are becoming increasingly important for generating high-quality, reproducible biological insights.
 DNA Synthesis
DNA Synthesis Vector Selection
Vector Selection Molecular Biology
Molecular Biology Oligo Synthesis
Oligo Synthesis RNA Synthesis
RNA Synthesis Variant Libraries
Variant Libraries Genome KO Library
Genome KO Library Oligo Pools
Oligo Pools Virus Packaging
Virus Packaging Gene Editing
Gene Editing Protein Expression
Protein Expression Antibody Services
Antibody Services Peptide Services
Peptide Services DNA Data Storage
DNA Data Storage Standard Oligo
Standard Oligo Standard Genome KO Libraries
Standard Genome KO Libraries Standard Genome Editing Plasmid
Standard Genome Editing Plasmid ProXpress
ProXpress Protein Products
Protein Products