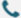The same drug can have a strong therapeutic effect on some individuals while showing limited efficacy in others. One of the key drivers behind this variability is genetic variation in the human genome, particularly single nucleotide polymorphisms (SNPs).
With the rapid development of SNP sequencing, SNP analysis, and advanced genotyping methods, researchers are now able to better understand disease susceptibility, drug response, and population genetics.
What Is SNP Genotyping?
Before exploring detection technologies, it is essential to understand what is SNP genotyping.
SNP genotyping refers to the process of identifying genetic variations at single nucleotide positions within the genome. A single nucleotide polymorphism (SNP) occurs when one base (A, T, C, or G) is altered, inserted, or deleted at a specific genomic location.
Following the completion of the Human Genome Project, SNP sequencing and SNP analysis have become critical tools in:
-
Disease risk assessment
-
Pharmacogenomics
-
Personalized medicine
-
Population genetics
The growing demand for these applications has driven rapid expansion in the SNP genotyping and analysis market, which is projected to reach billions of dollars globally in the coming years.
Overview of SNP Genotyping Methods
A wide range of genotyping methods are available for SNP detection, each offering distinct advantages in accuracy, throughput, and cost. The most commonly used approaches include SNP sequencing (such as Sanger and next-generation sequencing), qPCR-based SNP genotyping, DNA chip (microarray) detection, and emerging allele-specific PCR technologies, all of which play important roles in modern SNP analysis and genotyping workflows.
1. SNP Sequencing
SNP sequencing remains one of the most accurate approaches for SNP detection.
Sanger sequencing is considered the “gold standard” for SNP identification, with detection accuracy approaching 100%. It allows researchers to identify unknown SNP loci and precisely determine mutation types and positions.
In contrast, next-generation sequencing (NGS) enables high-throughput SNP analysis, allowing millions of SNPs to be detected simultaneously. These technologies are widely used in genome-wide association studies (GWAS) to identify genetic variants linked to complex traits.
Advanced targeted approaches such as parabon fx SNP capture AFDIL Mandeberg have also been applied in forensic genomics, enabling highly sensitive SNP profiling from challenging samples.
2. qPCR-Based SNP Genotyping
qPCR is one of the most widely used genotyping methods due to its speed, accuracy, and cost-effectiveness. It is especially popular in clinical diagnostics and targeted SNP analysis.
Among qPCR approaches, TaqMan probe-based assays are the most commonly used. These assays rely on fluorescence signals generated during amplification to distinguish between SNP alleles.
Other probe technologies used in qPCR-based SNP genotyping include MGB probes, which enhance specificity by stabilizing probe–target binding, and molecular beacon probes, which adopt a hairpin structure that enables flexible and highly selective hybridization to target sequences. While these probe designs improve the sensitivity and accuracy of SNP analysis, successful application still depends on careful probe design and optimization to ensure reliable and reproducible genotyping results.
3. Chip-Based SNP Detection (Microarray)
DNA chip technology is a powerful tool for large-scale SNP analysis and high-throughput genotyping. Microarrays enable thousands to millions of SNPs to be analyzed simultaneously by hybridizing labeled DNA samples to pre-designed probes immobilized on solid surfaces such as silicon or glass chips.
This approach is widely used in population-scale genotyping, disease association studies, and agricultural genomics. Chip-based platforms also play a significant role in the expanding SNP genotyping and analysis market, particularly for large cohort studies.
Choosing the Right SNP Genotyping Method
Each SNP detection approach offers unique benefits and limitations. Selecting the appropriate method depends on experimental goals, sample size, and required throughput.
Key evaluation criteria include design flexibility and success rate, allele detection accuracy, throughput and scalability, and cost per sample. In practice, combining multiple genotyping methods with robust SNP analysis pipelines often yields the most reliable results.
Conclusion
SNP genotyping technologies have become essential tools in modern genomics research. From traditional SNP sequencing to advanced qPCR and microarray platforms, each method contributes to more accurate and efficient genetic analysis.
As the SNP genotyping and analysis market continues to grow, innovations such as high-throughput sequencing and targeted capture technologies (e.g., parabon fx SNP capture AFDIL Mandeberg) are further expanding the possibilities of genomic research.
Understanding what is SNP genotyping and selecting the right genotyping methods will be key to advancing applications in precision medicine, drug development, and biotechnology.
About Us
Synbio Technologies is committed to providing professional and efficient DNA solutions for researchers worldwide. We offer advanced support for SNP sequencing, SNP analysis, and customized genotyping workflows to accelerate your research.
Related Services:
qPCR Probes
Oligo Pool Synthesis
Reference:
[1] Ayalew H, Tsang PW, Chu C, Wang J, Liu S, Chen C, Ma XF. Comparison of TaqMan, KASP and rhAmp SNP genotyping platforms in hexaploid wheat. PLoS One. 2019 May 22; 14(5): e0217222.[2] Li B, Liu Y, Hao X, Dong J, Chen L, Li H, Wu W, Liu Y, Wang J, Wang Y, Li P. Universal probe-based intermediate primer-triggered qPCR (UPIP-qPCR) for SNP genotyping. BMC Genomics. 2021 Nov 24; 22(1): 850.[3] S S, Fuke S, Nagasawa H, Tsukahara T. Single nucleotide recognition using a probes-on-carrier DNA chip. Biotechniques. 2019 Feb; 66(2): 73-78.
 DNA Synthesis
DNA Synthesis Vector Selection
Vector Selection Molecular Biology
Molecular Biology Oligo Synthesis
Oligo Synthesis RNA Synthesis
RNA Synthesis Variant Libraries
Variant Libraries Genome KO Library
Genome KO Library Oligo Pools
Oligo Pools Virus Packaging
Virus Packaging Gene Editing
Gene Editing Protein Expression
Protein Expression Antibody Services
Antibody Services Peptide Services
Peptide Services DNA Data Storage
DNA Data Storage Standard Oligo
Standard Oligo Standard Genome KO Libraries
Standard Genome KO Libraries Standard Genome Editing Plasmid
Standard Genome Editing Plasmid ProXpress
ProXpress Protein Products
Protein Products