The development of the CRISPR-Cas9 system has ushered in a new era of gene editing; which can edit single genes, polygenes, and even genomes in animals, plants, and microorganisms. The CRISPR-Cas9 system is commonly used in molecular breeding, drug discovery, and gene therapy. With convenient operation and low cost, CRISPR-Cas9 become one of the most popular gene editing technologies.
Off-Target Effects
The off-target phenomenon that occurs in the CRISPR-Cas9 system limits its further applications. sgRNA designed in CRISPR systems may recruit Cas9 nucleases to bind to unexpected regions and initiate cleavage, resulting in off-target effects. Ultimately, this leads to irreversible by-products after genome editing; including extensive mutations in the genome, and activation/inactivation of non-target genes that may lead to fatal or unwanted phenotypes, or activation of oncogenes that lead to cancer. However, if sgRNA binds to the target sequence, it will achieve powerful gene editing capabilities that produce expected results.
Off-Target Evaluations
Presently, the systematic summary and evaluation of on-target and off-target effects of sgRNA by researchers allow the CRISPR system to be utilized for more precise gene editing and gene therapy.
Various sequencing methods can be used to detect off-target sites, such as WGS (whole-genome sequencing), GUIDE-seq (genome-wide, unbiased identification of DSBs enabled by sequencing ), Digenome-seq (digested genome sequencing). There are also other algorithms/tools for sgRNA target search evaluation, and prediction of off-target effects, such as PEM-seq (prime-extension-mediated sequence), CRISPR-PLANT v2, CCTop (CRISPR/Cas9 target online predictor), etc.
Optimized CRISPR Systems
Major research institutions are continuously optimizing sgRNA design parameters and specific experimental needs hoping to achieve accurate gene editing. Computer-aided sgRNA design is one of the key steps for CRISPR gene editing. Currently, computational efforts are mainly focused on using computational models to improve the targeting efficiency of sgRNAs and reduce the off-target rate.
To date, several rather peculiar targeting rules have been established for Cas nucleases:
(1) Seed Region: Single mismatches within a PAM proximal seed region can completely disrupt interference, whereas PAM distal mismatches have much less of an effect.
(2) Mismatch Spread: When mismatches are outside the seed region, off-targets with spread out mismatches are targeted most strongly.
(3) Differential Binding versus Differential Cleavage: Binding is more tolerant to mismatches then cleavage.
(4) Specificity-Efficiency Decoupling: Weakened protein-DNA interactions can improve target selectivity while still maintaining efficiency.
CRISPR Services | Synbio Technologies
The rise in gene editing popularity spawned many suppliers but few companies have comprehensive service capabilities. As a synthetic biology enabling technology company, Synbio Technologies has an integrated DNA Reading (Sequencing), Writing (Synthesis), and Editing (Engineering) platform. With advanced design and leading manufacturing processes, we quickly deliver high-quality products to our customers.
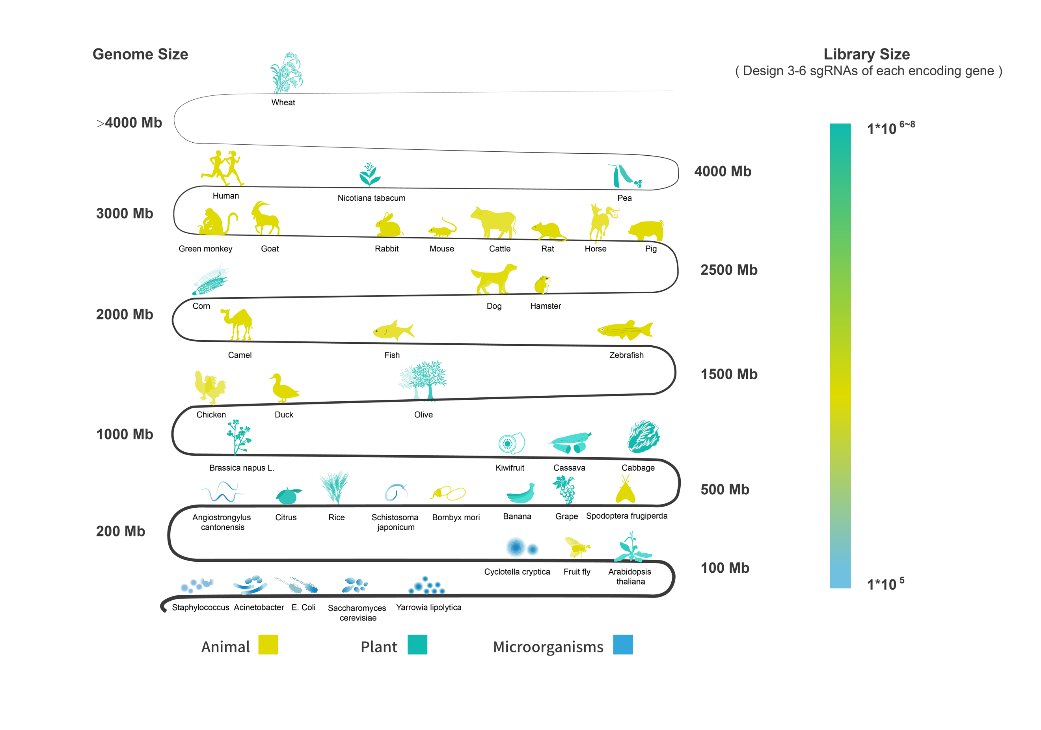
Relying on our proprietary design algorithm, Synbio Technologies’ experienced engineers can provide sgRNA design of single genes, specific pathways, and whole genomes for animals, plants, and microorganisms. This allows for efficient genome editing and high-throughput screening.
References
[1]Misha Klein, Behrouz Eslami-Mossallam, Dylan Gonzalez Arroyo, et al. Hybridization Kinetics Explains CRISPR-Cas Off-Targeting Rules. Cell Rep. 2018 Feb 6;22(6):1413-1423.[2]Hakim Manghwar, Bo Li, Xiao Ding, Amjad Hussain, et al. CRISPR/Cas Systems in Genome Editing: Methodologies and Tools for sgRNA Design, Off-Target Evaluation, and Strategies to Mitigate Off-Target Effects. Adv Sci (Weinh). 2020 Mar; 7(6): 1902312.

