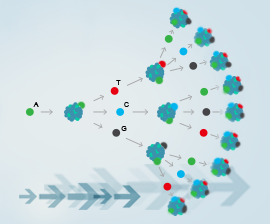
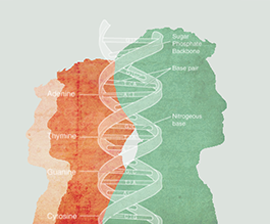
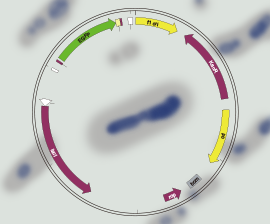
The CRISPR-Cas9 system is a kind of natural immune system that exists in bacteria. Whenever a virus invades, the system can edit foreign genomes to protect the immune functions of bacterial cells. Based on CRISPR-Cas9’s powerful genome editing capabilities, researchers have transformed the bacteria’s natural immune system into a gene editing tool that can be widely used in the laboratory. CRISPR-Cas9 is the third generation of gene editing tools following ZFN and TALEN.
Applications of CRISPR-Cas9
- Knock-out
Cas9 can cut the target genome and form double strand breaks of DNA. Under normal circumstances, cells will repair broken DNA using highly efficient non-homologous end joining (NHEJ). However, the mismatch of base insertion or deletion usually occurs in the process of repair, causing frame shift mutations, resulting in the target gene losing its function, thereby also enabling gene knockouts. In order to improve the specificity of the CRISPR system, one of the Cas9 domains can be mutated to form a Cas9 nickase nuclease that can only cut the single strand of DNA, causing the DNA gap. Therefore, two sgRNA sequences can be designed to form a double strand break, which targets the two complementary chains of DNA, so that two sgRNA specific binding target sequences can form a DNA fracture. Gene knockout is achieved through frame shift mutation during the repair process. - Knock-in
In the case that the DNA double-strand breaks, if the DNA repair template is entered into the cell, the fragment of the genome will be reconstructed by homologous recombination based on the repair template (HDR), thus enabling gene knock-in. The repair template is composed of the target genes and a homologous sequence (homologous arm) upstream and downstream of the target sequence that needs to be introduced. The length and position of the homologous arm are determined by the size of the editing sequence. The DNA repair template may be a linear/double-stranded deoxynucleotide chain or a double-stranded DNA plasmid. The incidence of HDR repair patterns in cells is low— usually less than 10%. In order to increase the success rate of gene knock-in, many scientists are currently working on improving HDR efficiency, synchronizing the edited cells to the most active cell division of HDR, promoting repair methods in HDR or using chemical methods to inhibit the gene NHEJ and improve the efficiency of HDR. - Repression or Activation
One important characteristic of Cas9 is that it can autonomously bind and cut the target gene by inactivating the two domains (RuvC- and HNH-) through point mutation. The formed dCas9 can only bind to the target gene under the mediation of sgRNA, so it does not have the function of cutting DNA. Therefore, if the dCas9 is combined with the transcription initiation site of the gene, the initiation of transcription can be blocked, thereby restraining gene expression. Binding the dCas9 to the promoter region of the gene can also integrate transcriptional inhibitors/activators to inhibit or activate the target gene transcription of the downstream target. Thus, the difference between dCas9 and Cas9 or Cas9 nickase is that the activation or inhibition caused by dCas9 is reversible and does not cause permanent changes to the genomic DNA. - Multiplex Editing
When multiple sgRNA plasmids are transferred into cells, they can edit multiple genes simultaneously and screen genome functions. In application, multiplex editing allows the use dual Cas9 nickases to improve the accuracy of gene knockout, wide-range genome deletions, and the ability to simultaneously edit different genes. Usually, two to seven different sgRNA can be constructed on one plasmid for multiple CRISPR gene editing. - Functional Genome Screening
When using CRISPR-Cas9 to edit genes, a large number of mutant cells may emerge. These mutant cells can help confirm whether the phenotypic changes are caused by genetic or genetic factors. The traditional method to screen genomics is to use shRNA technology. But shRNA has its limitations: it has a high rate of missing and can produce a false negative result because of its inability to inhibit all genes. The genomic screening function of the CRISPR-Cas9 system has the advantages of high specificity and irreversibility, so it has been widely used in genome screening. Presently, the genome screening function of CRISPR is used to screen the genes that can regulate the phenotype, such as the genes that are able to inhibit chemotherapeutic drugs or toxins, or genes affecting tumor migration. In addition, this function can be applied to the screening of potential genes by building virus screening libraries.