NGS: A “Magnifying Glass” for Tumor Recognition
Advancements in our understanding of tumor genetics have uncovered a remarkable truth: tumors are not just random growths, but rather genetic diseases fueled by intricate mutations. These mutations wield immense power, dictating the tumors’ creation, expansion, spread, and relentless progression. They can appear in various forms, from subtle alterations in DNA bases to the insertion or removal of DNA fragments. Sometimes, entire genes or their crucial parts can vanish. There are even fascinating rearrangements like inversions and translocations that occur. With this newfound knowledge, a wave of genetic testing initiatives has surged, seeking to detect these mutations and unlock crucial insights into tumor behavior.
Next-Generation Sequencing (NGS) has transformed oncology treatment by enabling a deeper understanding of the genomic landscape of cancer and facilitating personalized medicine approaches. NGS technology acts as a “magnifying glass,” pinpointing mutations at the molecular level.
The following applications highlight how NGS is changing oncology treatment by enabling precision medicine approaches that target the specific genetic alterations driving an individual’s cancer, enhancing treatment effectiveness and minimizing unwanted side effects.
- Tumor Profiling: NGS allows for the comprehensive profiling of tumor genomes, identifying genetic alterations such as somatic mutations, copy number variations, and structural variations. This information helps to characterize the specific molecular alterations driving a patient’s cancer and can guide treatment decisions.
- Companion Diagnostics: NGS-based tests can be developed as companion diagnostics to identify patients who are likely to respond to a particular targeted therapy. These tests assess the presence of specific biomarkers or mutations that predict response or resistance to a particular drug.
- Targeted Therapy Selection: NGS enables the identification of specific mutations or alterations in cancer-related genes that can be targeted with specific drugs. This information helps in selecting appropriate targeted therapies or clinical trials for patients based on the genomic profile of their tumor.
Explore More About Next-generation Sequencing
NGS refers to a high-throughput DNA sequencing technology that allows for rapid and cost-effective analysis of DNA or RNA samples. It is also known as Massively Parallel Sequencing or Second-Generation Sequencing. Unlike traditional Sanger sequencing, which can only sequence a single DNA fragment at a time, NGS techniques can simultaneously sequence millions to billions of short DNA fragments in parallel. NGS technology offers the ability to sequence the entirety of a genome or focus on specific gene regions, including sequencing all 22,000 coding genes (a whole exome) or just a small number of individual genes. What’s truly remarkable is that NGS methods now allow for the sequencing of an entire human genome in just one day, at a cost of approximately $1000.
NGS revolutionized the field of genomics by enabling researchers to sequence large amounts of genetic material in a short period of time. NGS platforms utilize various technologies and chemistries to achieve sequencing. The most commonly used NGS platforms is Illumina’s sequencing-by-synthesis technology.
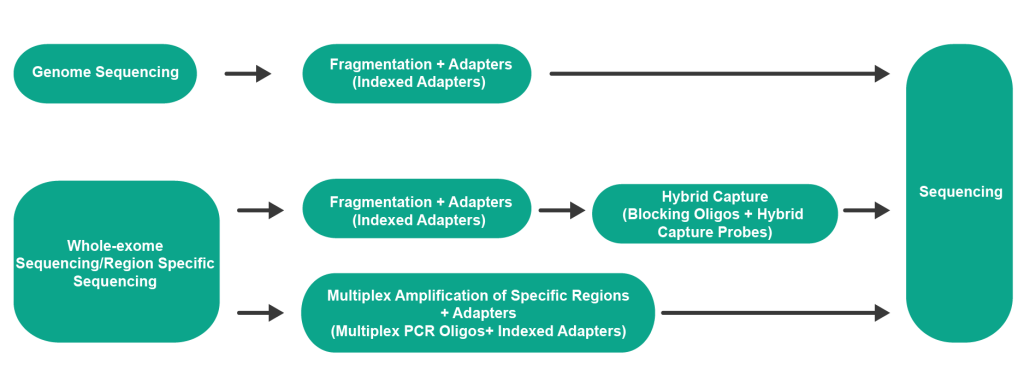
The NGS workflow typically involves the following steps:
- Sample Preparation
- Library Construction: DNA or RNA samples are fragmented, and adapters are added to the fragments to facilitate sequencing.
- Sequencing: The prepared library is loaded onto an NGS platform, generating detailed DNA sequence data.
- Data Analysis
Throughout the NGS process, four types of oligos are used, including indexed adapters, multiplex PCR oligos, hybrid capture probes, and blocking oligos.
- Indexed Adapters: Indexed adapters link the DNA of the sample to the sequencer, which is essentially a short sequence of bases, typically 15-80nt in length.
- Multiplex PCR Oligos: Multiplex PCR is a PCR technique that amplifies multiple DNA sequences simultaneously. In multiplex PCR, several different primers can be used so that multiple target sequences are amplified in the same reaction.
- Hybrid Capture Probes: According to the principle of DNA base complementary pairing, the probe sequence is complementary to the target sequence and has a biotin marker.
- Blocking Oligos: Blocking primers prevent hybridization between the capture probe and the junction region of the library in capture sequencing, reducing the off-target effect of the capture probe and improving the specificity of capture sequencing.
Synbio Technologies | NGS Oligos
Ensure optimal performance by leveraging Synbio Technologies‘ selection of indexed adapters, capture probes, and blocking oligos to minimize external contamination or splicing errors in your sequence results.
|
NGS Oligo Type |
Length (bp) |
Turnaround Time |
Deliverables |
|
Indexed Adapters |
60-70 |
5-7 |
Products in Tube/Plate/Chip, Comprehensive QC Report (HPLC, MS) |
|
Capture Probes
|
80-120 |
Ask us |
|
|
Multiplex PCR Oligos |
20-60 |
Ask us |
|
|
Blocking Oligos |
20-80 |
5-7 |
|
|
Other NGS Oligos |
10-150 |
5-7 |
* If you want to learn more, please contact our team.
Why choose us?
- High Quality: ISO9001 & ISO13485 quality management system standards, strict process control, and QC testing.
- Low Cross Contamination: Strict purification standards, with a maximum purity of over 95% and low external interference rate.
- Customization: Various purification methods, several synthesis specifications, diversified shipping forms, and customized services.
References
- Shirdarreh M, Aziza O, Pezo R C, et al. Patients’ and oncologists’ knowledge and expectations regarding tumor multigene next-generation sequencing: A narrative review[J]. The Oncologist, 2021, 26(8): e1359-e1371.
- Denny JC, Collins FS. Precision medicine in 2030-seven ways to transform healthcare. Cell. 2021 Mar 18;184(6):1415-1419.
- Wakai T, Prasoon P, Hirose Y, et al. Next-generation sequencing-based clinical sequencing: toward precision medicine in solid tumors[J]. International journal of clinical oncology, 2019, 24: 115-122.
- Vincent A T, Derome N, Boyle B, et al. Next-generation sequencing (NGS) in the microbiological world: How to make the most of your money[J]. Journal of microbiological methods, 2017, 138: 60-71.
- Muzzey D, Evans EA, Lieber C. Understanding the Basics of NGS: From Mechanism to Variant Calling. Curr Genet Med Rep. 2015;3(4):158-165.
- Behjati S, Tarpey PS. What is next generation sequencing? Arch Dis Child Educ Pract Ed. 2013 Dec;98(6):236-8.