Infectious diseases have been around since the dawn of time. Currently, there are two ways to test for infectious diseases: testing for pathogens or testing for antibodies that the body produces to fight the pathogens. Pathogen detection can be performed via antigen detection or nucleic acid detection. Following the worldwide outbreak of COVID-19, nucleic acid detection techniques, based on a molecular level, have gradually become the main method for pathogen detection. Let’s breakdown the mainstream technologies currently used for pathogen detection.
Relative Quantification in Real-time (RT–PCR)
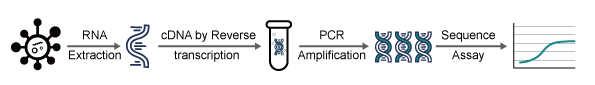
In RT-PCR, isolated viral RNA is reverse transcribed into cDNA, which is subsequently amplified by PCR and identified by a fluorescent probe. One can tell whether a sample includes the viral sequence by looking at the fluorescent signal in the amplified product. Relative quantification in real-time (RT–PCR) is a rapid and sensitive method to detect low amounts of pathogen and offers important physiological insights on mRNA expression levels. It is also the primary method for early detection of the spread of infectious diseases.
Isothermal Reverse Transcription Amplification
Isothermal reverse transcription amplification is a technique for rapid amplification of nucleic acids at a constant temperature by adding enzymes with different activities and their corresponding primers. In recent years, isothermal reverse transcription amplification has advanced significantly. It exhibits considerable superiority in field diagnostics because it no longer depends on a thermal cycler. Currently, common isothermal signal amplification techniques include loop-mediated isothermal amplification (LAMP), strand displacement amplification (SDA), rolling circle amplification (RCA), and nucleic acid sequence-based amplification (NASBA). LAMP is highly specific, distinguishing up to eight specific locations of the DNA or RNA template, and is the most mature isothermal amplification technique among others. NASBA is a simple isothermal signal amplification method that is widely used in monitoring and controlling the transmission of animal diseases.
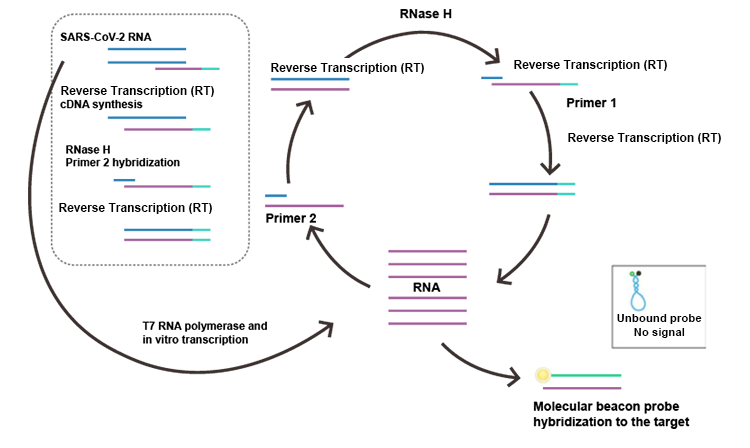
Detection of SARS-CoV-2 Virus by Real-time NASBA
NGS Sequencing
Nucleic acid sequences can only be obtained through gene sequencing. NGS is a DNA sequencing technology based on PCR and gene chips. The basic NGS sequencing process involves fragmenting DNA/RNA into multiple pieces, adding adapters, sequencing the libraries, and reassembling them to form a genomic sequence. By collecting fluorescent labeling signal or chemical reaction signal, we can
obtain gene sequences. NGS can complete the sequencing of several hundred thousand to several million sequences at one time and enables de novo sequencing of unknown viruses at the genome level to obtain the gene sequences of pathogens. Compared with RT-PCR, NGS can detect a variety of viral sequences while revealing the exact sequence changes, making it a typing information, and is a powerful tool for virus detection, genotyping, and tracing.
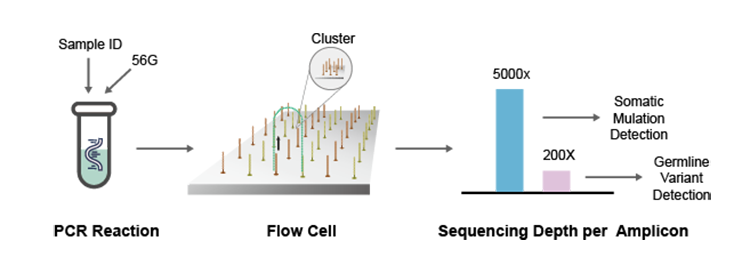
NGS Sequencing Principle
CRISPR-Cas System-based Detection
More recent applications of CRISPR for the development of infectious disease diagnostics take advantage of collateral cleavage induced by Cas12a and Cas13a nucleases. Under the guidance of sgRNA, a crRNA-Cas12a or crRNA-Cas13a complex binds and cleaves target nucleic acid with high specificity. When the crRNA-Cas13a or crRNA-Cas12a complex contacts the target nucleic acid, non-target RNA connected to a fluorescent reporter is also cleaved, producing a fluorescent signal for pathogen identification. Compared with isothermal amplification, CRISPR-Cas system detection uses isothermal amplification in combination with Cas enzyme trans-cutting. Quick on-site detection is made possible by the identification of sgRNA, which also improves specificity following isothermal amplification. CRISPR-Cas system-based detection is a next-generation molecular diagnostic technology that is expected to disrupt the field of in vitro diagnostics in the near future.
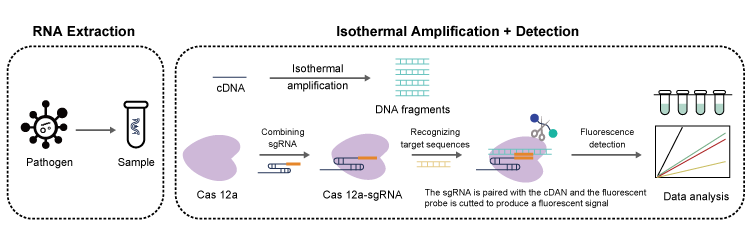
Workflow | Infectious Disease Diagnosis based on CRISPR-Cas System Detection
Selecting the Appropriate Detection Method
| Detection Method | Advantages | Limitations | Applications | Detection Duration |
|---|---|---|---|---|
| RT-PCR |
|
|
Can be used to detect a variety of viruses (e.g., adenovirus, rotavirus, astrovirus & enterovirus) | 2 – 4 h |
| Isothermal Reverse Transcription Amplification |
|
|
Can be used to detect Mycobacterium tuberculosis, Mycobacterium trachomatis, HIV, and Severe Acute Respiratory Syndrome viruses | 30 – 60 min |
| NGS Sequencing |
|
|
Widely used for diagnosis of viral respiratory pathogens (e,g. Mycobacterium tuberculosis, Pseudomonas aeruginosa strains, and Staphylococcus aureus strains) | 24 h + |
| CRISPR-Cas System-based Detection |
|
|
Cas12a for accurate genotyping of HPV viruses, Cas13a for detection of dengue and Zika viruses, & Cas14a for single nucleotide polymorphism (SNP) detection | 45 – 60 min |
Synbio Technologies Specializes in Diagnostic Probe and Oligo Synthesis
No matter which detection method you choose, you will need primers and probes as your core material. Synbio Technologies provides a one-stop solution for in vitro diagnostics with cutting-edge DNA manufacturing, precise probe design, and a fast delivery cycle. High-quality primers and probes, synthesized by Synbio Technologies, can greatly improve experimental efficiency and detection accuracy in your research. Our diagnostic probes such as TaqMan/MGB qPCR probes, Molecular Beacons, Scorpion Probes, LNA Probes, Dual Quencher Probes, and RAA/RPA Probes can be used in molecular hybridization and PCR. We also offer customized services to meet the various application requirements in the field of molecular diagnostics.
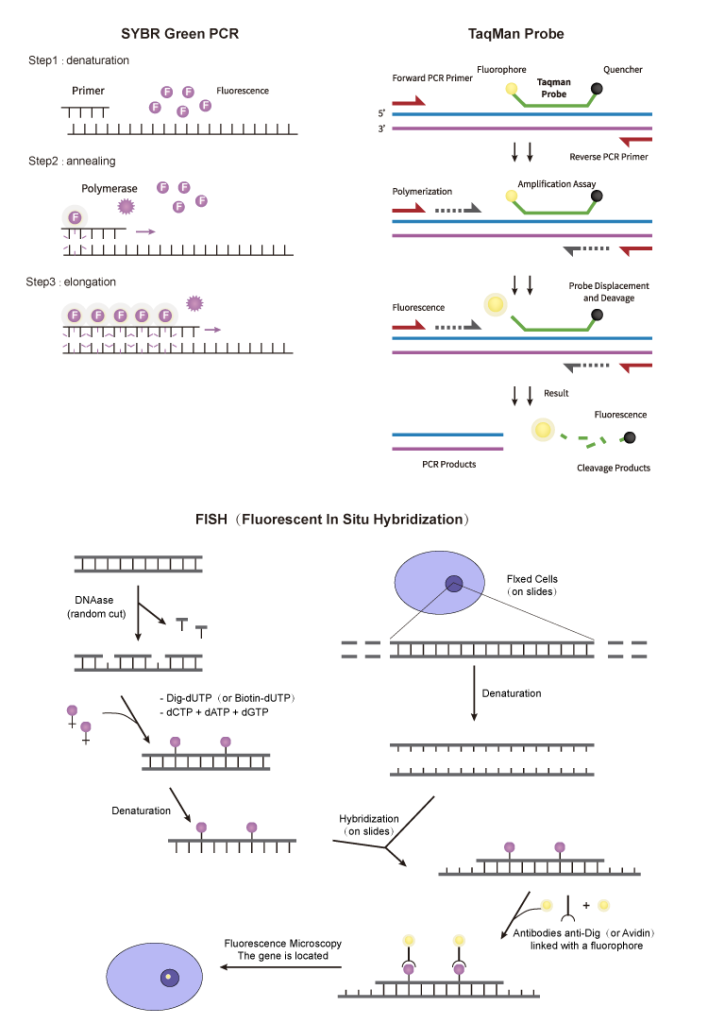
Probe Types
Case Study
Oligonucleotide synthesis refers to the chemical synthesis of single-chain, short fragments via solid-phase phosphite amides. The synthesis of each base requires four steps: deprotection, activation and coupling, acetyl closure, and oxidation stability. Due to efficiency problems with the chemical reaction itself, the crude products cannot reach 100% purity. In addition to the target sequence, short error fragments and impurities will also be generated during the synthesis process, which will affect the experimental results as well as the progress of the project.
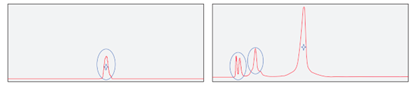
Conventional probe purification (left) vs. Synbio Technologies purification (right)
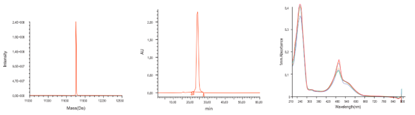

Quality Control Results
Synbio Technologies upgraded the purification process and optimized the ratio of reagents. We can achieve up to 98% purity in target sequence separation.
The high quality control results from Synbio Technologies show that our probes are prepared with theoretical molecular weights matching those of the samples being tested, no signs for discharges or peaks in NMR spectra. In this case, our probe products were purified better than the market average. When you need customized, oligos & probes, with high-purity – Synbio Technologies has you covered!
References:
- Hu WS, Hughes SH. HIV-1 Reverse T Cold Spring Harb Perspect Med. 2012 Oct 1;2(10):a006882. doi: 10.1101/cshperspect.
- Sun Y, Yu L, Liu C, Ye S, Chen W, Li D, Huang W. One-tube SARS-CoV-2 Detection Platform Based on RT-RPA and CRISPR/Cas12a. J Transl Med. 2021 Feb 16;19(1):74. doi: 10.1186/s12967-021-02741-5.
- Kia V, Tafti A, Paryan M, Mohammadi-Yeganeh S. Evaluation of Real-time NASBA Assay for the Detection of SARS-CoV-2 Compared with Real-time PCR. Ir J Med Sci. 2022 Jun 6;1–7. doi: 10.1007/s11845-022-03046-2.
- Deurenberg RH, Bathoorn E, Chlebowicz MA, etc. Application of Next Generation Sequencing in Clinical Microbiology and Infection Prevention. J Biotechnol. 2017 Feb 10;243:16-24. doi: 10.1016/j.
- Soroka M, Wasowicz B, Rymaszewska A. Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR? Cells. 2021 Jul 29;10(8):1931. doi: 10.3390/cells10081931.
- Nowrousian M. Next-generation Sequencing Techniques for Eukaryotic Microorganisms: Sequencing-based Solutions to Biological Problems. Eukaryot Cell. 2010 Sep;9(9):1300-10. doi: 10.1128/EC.00123-10.
- Ortiz-Cartagena C, Fernández-García L, Blasco L, etc. Reverse Transcription-Loop-Mediated Isothermal Amplification-CRISPR-Cas13a Technology as a Promising Diagnostic Tool for SARS-CoV-2. Microbiol Spectr. 2022 Sep 28:e0239822. doi: 10.1128/spectrum.02398-22.