Over the past few decades, cancer immunotherapy has dramatically altered approaches toward oncology therapy. Although treatment methods such as checkpoint blocking can produce significant clinical responses, the efficacy is uneven between cancer patients and different cancer types. Due to the high heterogeneity and complexity of the tumor microenvironment, the infiltrating lymphocytes need to be analyzed in order to characterize the basic properties of different types on immune cells.
T Cells, also known as the T lymphocytes, play an important role in cell-mediated immune regulation. T cells can be separated from other lymphocytes (e.g. B cells and natural killer cells) by the T Cell receptors on the surface of the cell. Most of the human T-cells are the αβT cell since the α and β- chains are reformed on the receptors. To better understand T cell’s involvement in this regulation, Synbio Technologies provides T-cell receptors library sequencing and bioinformatics analysis. By using the targeted amplification technique, this analysis obtains the CRR3 sequence of the TCR α&β chains. Synbio Technologies also provides high throughput sequencing and bioinformatics to analyze the T-cell acceptor library. This analysis provides sensitive, precise, and reliable data for the clinical trial and innovation research related to cancer, infection or autoimmunity.
Featured Advantages
- Unique in-house data mining tools to pre-process antibody sequencing to minimize sequencing errors such as false positives
- Experienced bioinformatics analysis research team capable of obtaining highly valuable antibody information
Synbio Technologies Case Study
Based on the clients’ original sample, Synbio Technologies extracted the RNA, and established the T-cell library by RACE amplification. Synbio Technologies provided an intact T-cell library analysis report by high throughput sequence and bioinformatics analysis.
Results:
1. Comparison with VDJ gene reference sequences
Synbio Technologies comprised the sequenced resulted with the reference sequence from data base to remove the invalid sequence, including unmatched and stop codon region. Then decide the VDJ expression condition and specific CFR3 region by the comparison results.
2. Frequency of VDJ gene usage analysis
2. Frequency of VDJ gene usage analysis
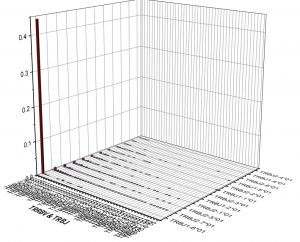
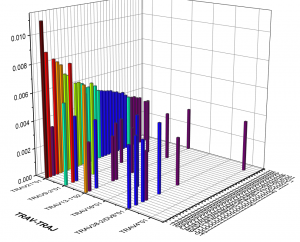

3. Analysis of CDR3 region length
4. Analysis of CDR3
5. Ordering of total length sequence
| Amino Acid Gene Length | Amino Acid Length | Abundance |
|---|---|---|
| 1491 | 147 | 0.096324 |
| 620 | 139 | 0.044324 |
| 540 | 143 | 0.040395 |
| 516 | 147 | 0.041955 |
| 496 | 141 | 0.043976 |
| 433 | 145 | 0.041868 |
| 366 | 109 | 0.041762 |
| 387 | 148 | 0.039055 |
| 504 | 143 | 0.042773 |
| 260 | 148 | 0.032298 |
6. Saturation Analysis of sequencing results
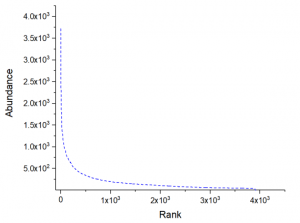
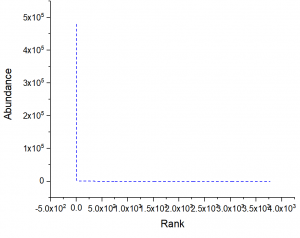
7. The difference of the same combination of V gene and J gene combination in two samples
Service Specifications
| Service | Detail | Analysis | TAT (Business Days) | Deliverable |
|---|---|---|---|---|
| T-cell antibodies sequence Service | Data mining of sequencing, Antibody information analysis etc. | R, Python Software etc. | 7-10 | Analysis report |

